In our previous blog, we covered the highlights of our Advanced Report Customization in VSClinical webcast in the context of germline clinical reports. Now, we bring you the next of the series: somatic clinical reports. In the recent webcast, Advanced Report Customization, we covered a range of somatic-focused clinical reports, demonstrating how easy it is to create AMP guideline-based clinical… Read more »
The articles we saw this November and December cited a wide range of applications of our product suite. The following publications feature usage of our SNP & Variation Suite, VSClinical, and VarSeq products. We see them being utilized to identify loci associated with facial eczema in New Zealand sheep, somatic mutation response impact, and assisting in estimating breast cancer risk… Read more »
As the number of genes on a gene panel increases, there is the possibility of picking up variants of medical significance that are not related to the primary indication for the test. Especially with large gene panels, exomes, and genomes, it is medically and ethically important to report variants that may be actionable to the patient. These include variants implicating… Read more »
When it comes to clinical variant analysis for germline variants using ACMG guidelines, we understand that the clinical report is essentially the receipt for your services for the patient. In our recent webcast, Advanced Report Customization in VSClinical, we displayed several examples of report templates to show off the range of possibilities our users have to format their clinical report…. Read more »
In February 2020, the American College of Medical Genetics (ACMG) and the Clinical Genome Resource (ClinGen) published a joint consensus on standards for the interpretation and reporting of copy number variants (CNVs) ranging from large CNVs spanning multiple genes to small intragenic events1. The guidelines consist of over 80 different criteria which are arranged into five distinct sections. These extensive… Read more »
Thank you to everyone who attended Using Golden Helix CancerKB to Accelerate NGS Cancer Testing! I had a great time showing off Golden Helix CancerKB and how it can enhance NGS analysis in cancer, I certainly hope you enjoyed it! If you were unable to make the live session, we have added the on-demand recording to our site, or we… Read more »
As a newer member of the Golden Helix FAS Team I, like many of our customers, went through a phase of learning the ins, outs, and shortcuts of clicking around the program. In an effort to expedite new users coming up to speed, or to teach some tricks to experienced users, I’ve compiled a list of the top things I… Read more »
Recently we have released blog posts discussing updates to annotations in VarSeq such as ClinVar and COSMIC. Keeping with that trend, in this blog post, I will discuss the most recent updates to the Golden Helix CancerKB database. For those how may be unfamiliar with the Golden Helix CancerKB source, it is a professional curated set of interpretations for the… Read more »
In order to thoroughly assess a variant’s pathogenicity, it is important to take into account the variant’s effect on splicing. While the interpretation of variants that disrupt the pairs of bases at the beginning of a splice site is fairly straightforward, variants resulting in the introduction of a novel splice site are more difficult to interpret. In this blog post,… Read more »
With the release of VarSeq 2.2.4 just around the corner, I want to detail some new 2.2.4 features that will enhance somatic variant annotation and fusion analysis within VSClinical. A webcast back in July showed some of these updates in action, so if you are looking for some more content on this topic, I would highly recommend checking out the… Read more »
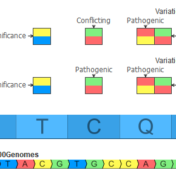
In the September 2021 monthly update to our curated ClinVar track, we made some changes that will result in roughly another 7,000 Likely Pathogenic and Pathogenic variants being available for annotation and use in the ACMG auto-classification system. Consensus Between Labs ClinVar has nearly one million unique variant classification records that are curated into multiple annotation tracks used in VarSeq and VSClinical on a monthly basis. Clinical… Read more »
Welcome to the August edition of our customer publications blog post! Each month we spotlight a few recently published articles by our incredible Golden Helix customers. With users spanning both research and clinical spaces, the topics vary widely across many fields. This month, we will be highlighting VSClinical users and the guided workflow. Host Genetics and Antiviral Immune Responses in… Read more »
Clinical labs often maintain gene panels, which are lists of genes with evidence of disease association. These panels are used to prioritize variants and limit interpretations to a predefined set of test-specific genes. In general, gene panels should be stored independently of any specific project or interpretation, as it is common for an individual gene panel to be generally applicable… Read more »
This blog post will cover an exciting new VSClinical feature in the upcoming VarSeq release. The ACMG Previously Interpreted Variants feature allows users to integrate databases of expert-curated variant interpretations into their VSClinical workflows. These data sources store variant-level interpretation data, including the classification, associated disorders, interpretation text, and scored criteria for each variant, along with notes providing a justification… Read more »
One of the many tricks of encoding so much functionality into so little space in eukaryotic genomes is the ability to produce multiple distinct mRNAs (transcripts) from a single gene. While one transcript is often the dominant one for a given tissue or cell type, there are, of course, exceptions in the messy reality of biology. It doesn’t take many… Read more »
Thank you to those who attended the recent webcast, “Exome Analysis with VS-CNV & VSClinical: Updated Strategies & Expanded Capabilities”. For those who could not attend but wish to watch, here is a link to the recording. In this webcast, we covered the capabilities and updates that have been incorporated into VarSeq that enhance whole exome sequencing workflows. The new… Read more »
The GenomeAsia 100K Variant Frequency database is a pilot annotation source now available to our users. This valuable database offers a deep characterization of specific populations in Asia that can be used to drive genetic studies. GenomeAsia is comprised of whole-genome sequencing data of over 1,000 individuals from 219 populations across Asia. Using this as an annotation, users can analyze… Read more »
Orphanet is a public database available in VarSeq that aims to improve the current understanding and treatment options for patients affected by rare diseases. This resource was established in France in 1997 and has gradually grown to cover a consortium of over 40 countries in Europe. The primary goal of this database is to provide a universal nomenclature to classify… Read more »
Clinical testing labs produce reports as the end product of the NGS variant detection and interpretation workflow. Necessarily, the content, detail, and presentation of the report needs to be specialized to each clinical lab, and potentially each offered test. Our last blog post introduced the new Word-based report templates in VSClinical. In this blog post, we will introduce and explore… Read more »
This morning I released a new version of my eBook “Clinical Variant Analysis – Second Edition.” The clinical interpretation of variants in Next-Gen Sequencing is a quickly evolving field. While the body of knowledge is growing exponentially, experts have to derive sound, clinical decisions leveraging an ever-expanding set of specialty databases, clinical publications, and algorithms that are designed to predict the… Read more »