September 27, 2017 12:00 PM, EDT While Copy Number Variants are important to detect and interpret in many clinical genetic tests, labs have been without a comprehensive solution that integrates the annotating and reporting of high-quality CNV alongside their existing NGS variants. Golden Helix has developed and validated with our clinical partners a specialized NGS-based CNV caller capable of detecting… Read more »
Annotating with gnomAD: Frequencies from 123,136 Exomes and 15,496 Genomes When the Broad Institute team lead by Dan MacArthur announced at ASHG 2016 that the successor to the popular ExAC project (frequencies of 61,486 exomes) was live at http://gnomad.broadinstitute.org/, I thought their servers would have a melt-down as everyone immediately jumped on and started looking up their favorite genes and… Read more »
There is no doubt that we have big data in the field of genomics in general and Next Generation Sequencing specifically. Illumina’s latest HiSeq X can produce 16 genomes per run, resulting in terabytes of raw data to crunch through. Yet all that crunching is not the hard part. So, what is the main obstacle to scientists being able to… Read more »
Probably one our most popular public annotation sources we curate and update is the database of Non-Synonymous Functional Predictions (dbNSFP). In it’s recent update, it has expanded the predictions to include FATHMM-MKL and VarSeq now incorporates this new prediction into its voting algorithm of now 6 different discrete predictions per variant. You can update to dbNSFP 3.0 using the built-in… Read more »
As VarSeq’s adoption has grown among analysts using whole exome data to diagnose rare diseases, a couple of family designs outside of the common trio of an affected child and both parents have come up frequently. While having both parents provides the maximum power to discover de novo mutations and recessively inherited variants, it is not always possible to contact… Read more »
Yesterday, our VP of Product and Engineering, Gabe Rudy, presented VSReports to the Golden Helix community for the first time in a live webcast; Authoring Clinical Reports in VarSeq. It was an excellent presentation. Gabe highlighted VSReports’ ability to take the output of tertiary analysis to a customized clinical grade report in one click. He also gave an overview of… Read more »
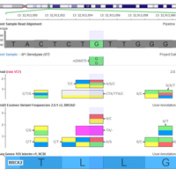
Have you ever scratched your head when looking up a variant and it seems like the number you have for its position is one off from what it looks like in the file or database? You may be running into the dreaded world of 1-based versus 0-based coordinate representation! If it’s any consolation, I can promise that all the bioinformaticians… Read more »
This last week I had the pleasure of attending the fourth annual Clinical Genome Conference (TCGC) in Japantown, San Francisco and kicking off the conference by teaching a short course on Personal Genomics Variant Analysis and Interpretation. Some highlights of the conference from my perspective: Talking about clinical genomics is no longer a wonder-fest of individual case studies, but a… Read more »
Yes, I said it. “Them be fighting words” you may say. Well, it’s worth putting a stake in the ground when you have worked hard to have a claim worth staking. We have explored the landscape, surveyed the ravines and dangerous cliffs, laboriously removed the boulders and even dynamited a few tree stumps. Stake planted. Ok, so now I’m going… Read more »