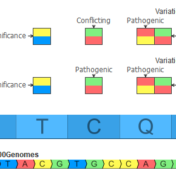
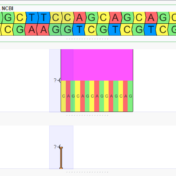
In the September 2021 monthly update to our curated ClinVar track, we made some changes that will result in roughly another 7,000 Likely Pathogenic and Pathogenic variants being available for annotation and use in the ACMG auto-classification system. Consensus Between Labs ClinVar has nearly one million unique variant classification records that are curated into multiple annotation tracks used in VarSeq and VSClinical on a monthly basis. Clinical… Read more »
Merging variant records, VCFs, across samples is important when performing trio or family analysis as it ensures that hereditary relationships can be properly inferred. There are many ways to represent a single variant. Insertions and deletions may be right or left aligned, prefixes and suffixes can be added, and adjacent variants in the same sample may be combined or split… Read more »
Clinical labs often maintain gene panels, which are lists of genes with evidence of disease association. These panels are used to prioritize variants and limit interpretations to a predefined set of test-specific genes. In general, gene panels should be stored independently of any specific project or interpretation, as it is common for an individual gene panel to be generally applicable… Read more »
In this month’s Customer Publication blog, I will highlight four studies that provided further insights into conditions that are typically identified in early childhood. As you will see as you read the summaries of each publication, both Golden Helix software platforms (VarSeq and SNP & Variation Suite or SVS for short) were instrumental in exploring the genetic factors that influence… Read more »
This blog post will cover an exciting new VSClinical feature in the upcoming VarSeq release. The ACMG Previously Interpreted Variants feature allows users to integrate databases of expert-curated variant interpretations into their VSClinical workflows. These data sources store variant-level interpretation data, including the classification, associated disorders, interpretation text, and scored criteria for each variant, along with notes providing a justification… Read more »
One of the many tricks of encoding so much functionality into so little space in eukaryotic genomes is the ability to produce multiple distinct mRNAs (transcripts) from a single gene. While one transcript is often the dominant one for a given tissue or cell type, there are, of course, exceptions in the messy reality of biology. It doesn’t take many… Read more »
Next-gen sequencing (NGS) comprises many sophisticated steps that are often compressed into three major sections: library prep, sequencing, and data analysis. Obviously, the goal is to simplify each of these steps, but more often than not, there is a need for multiple tools to complete each one. Regarding the data analysis, Golden Helix seeks to provide simple yet comprehensive solutions… Read more »
VSPipeline is becoming a very popular tool among VarSeq users as it is essential for creating repeatable clinical workflows that can be executed in automated fashion. Since VSPipeline is a command-line tool, I think it would be helpful to discuss some of the best practices along with helpful tips for getting the most out of VSPipeline. Some of you may be less familiar with VSPipeline, so I want to cover how to set up the first run along with sharing the helpful tips as they arise. … Read more »
One of the inherent realities of next-generation sequencing is the ongoing updates to the human reference genome—one of the strongest recommendations to take the original sequencing data and remap to the latest genome assembly. However, there are several reasons why remapping may be impractical. So, an alternative solution is needed to convert the data running through an initial mapping to… Read more »
A common feature request from Golden Helix customers is to curate and make available genome assemblies for different plant and animal species. These requests commonly come from SVS users as many research projects are being carried out, and having the genome assembly available for analysis is essential. That being said, Golden Helix has an SVS Tutorial available that walks users… Read more »
Golden Helix has just released VarSeq v2.2.3. In this update, there are notable changes that can improve CNV calling capabilities covered in this webcast. The topics discussed included: Accounting for GC content Improvements to CNV quality flags Using target filtering Updates to the CNV sensitivity and precision settings This blog post will elaborate on these capabilities and demonstrate how they… Read more »
Thank you to those who attended the recent webcast, “High Precision Exome CNV Detection with VS-CNV”. For those who could not attend but wish to watch, here is a link to the recording. This webcast delved into the complex world of CNV calling for whole-exome samples, which presents unique challenges that require specific considerations and strategies. Over the past several months,… Read more »
I want to take this opportunity to highlight and briefly discuss some of the key features and updates that have been incorporated into VarSeq 2.2.3. Some of you may have attended the webcast that covered the prominent new features added to VarSeq, which are the updates to improve whole-exome analysis workflows, namely improved CNV calling in whole-exome datasets. However, there… Read more »
With the latest release of VarSeq, we have made significant updates to our handling of the interaction of variants and genes. This includes the support for non-coding transcripts, improved splice site predictions, and updates to gene and transcript annotations. We received several questions regarding how decisions are made in the software regarding genes and transcripts with these gene-related changes. This… Read more »
While VarSeq has always had excellent support for variant interpretation and analysis, we continue to find new edge cases in the clinical literature that improve our interpretation capabilities. In this blog, we will be covering some of the new improvements in VarSeq to support the interpretation of non-coding and splice site variants. Transcript Annotation Improvements Let’s start by covering some… Read more »
Thank you to those who attended the recent webcast, “Exome Analysis with VS-CNV & VSClinical: Updated Strategies & Expanded Capabilities”. For those who could not attend but wish to watch, here is a link to the recording. In this webcast, we covered the capabilities and updates that have been incorporated into VarSeq that enhance whole exome sequencing workflows. The new… Read more »
In this blog, we will be covering new assessment catalogs and how they work to improve saving and tracking variant interpretations. VarSeq is a variant analysis tool that effectively analyzes single nucleotide (SNVs) and copy number variants (CNVs) in both cancer and germline workflows. Because VarSeq enables such diverse variant analysis, there are many research labs and institutions that evaluate… Read more »
Thank you to those who attended the recent webcast, “VSWarehouse: Tracking Changing Variant Evidence and Classifications”. For those who could not attend but wish to watch, here is a link to the recording. The webcast covered some general highlights of VSWarehouse value but also presented some specific capabilities covering the ClinVar classification tracker. Golden Helix provides complete solutions to handle… Read more »
The recent release of VarSeq 2.2.2 brings our Word report template system, previously featured in VSClinical AMP, to the VSClinical ACMG workflow. This blog post will describe how to use the Word template system using one of our shipped templates as well as how to start customizing your own templates. We will cover the three different report templates that ship… Read more »
Our latest VarSeq release is one of the largest we’ve ever had, boasting an extensive list of new features and improvements. As part of this release, we have dramatically expanded our support for splice site analysis. This includes improvements to our novel splice site algorithm and support for splice site effect prediction along with several other small improvements. Novel Splice… Read more »