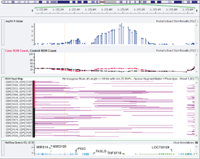
I spent a very eventful week at the Molecular TriCon in downtown San Francisco, and have been pondering the very clear trends that emerged by attending the clinical and NGS focused talks. Cancer gene panels make sense economically and as “massively parallel” tests to inform therapy, but they are bound to get more complex. Liquid biopsies of circulating tumor DNA… Read more »
TriCon 2015 was well worth the visit to San Francisco. The combination of extensive programming in conjunction with a large exhibition makes it a must-attend event for scientist and professionals in our industry and the conference seems to grow year after year. This year, we paid a lot of attention to the Clinical Sequencing portion of the event. In this track,… Read more »
A few of our customers have published recently and I would like to take the time to both recognize them for their achievement and pass on their articles. Enjoy! Kazima Bulayeva at the Vavilov Institute of General Genetics and her colleagues recently published Genomic Structural variants are linked with intellectual disability in the Journal of Neural Transmission. The paper looks at mutations… Read more »
Last month, Dr. Bryce Christensen presented Population-Based DNA Variant Analysis via webcast. The webcast reviewed the fundamentals of population-based variant analysis and demonstrated some of the tools available in SVS for analysis of both common and rare variants such as the SKAT-O method, as well as other functions for annotation, visualization, quality control and statistical analysis of DNA sequence variants. Here… Read more »
Today, Golden Helix and Fluxion Biosciences announced a collaboration in a value-added reseller relationship. The relationship will bring the VarSeq software application to Fluxion’s global client base providing them with a method to study tumor DNA in the circulation. Fluxion is proud to offer the capability as it helps move them toward their goal of offering a complete sample-to-answer workflow for… Read more »
Our Genomic Prediction webcast in December discussed using Bayes-C pi and Genomic Best Linear Unbiased Predictors (GBLUP) to predict phenotypic traits from genotypes in order to identify the plants or animals with the best breeding potential for desirable traits. The webcast generated a lot of good questions as our webcasts generally do. I decided to begin to share these Q&A… Read more »
In just 5 days, the 22nd International Molecular Medicine Tri-Conference (Tri-Con) will kick off in San Francisco. This year, Tri-Con will offer over 3,000 attendees 6 symposia, over 20 short courses, and 17 conference programs focused on drug discovery, genomics, diagnostics, and information technology surrounding, molecular medicine. Both Dr. Andreas Scherer, CEO of Golden Helix and Gabe Rudy, our Vice President of… Read more »
Today, we at Golden Helix announced our collaboration with PreventionGenetics as they prepare to implement the VarSeq software into their exome sequencing pipeline. The VarSeq software will allow PreventionGenetics to offer an exome test by dramatically speeding up the analysis process. VarSeq will narrow down sequence data into gene(s) of interest based on inheritance patterns, facilitating the identification of clinically relevant… Read more »
Genotype imputation is a statistical technique for estimating sample genotypes at loci that were not directly assayed by sequencing or microarray experiments. There are several reasons why you might want to use imputation in a research study. For example: Improve call rates in GWAS by imputing sporadic missing genotypes Harmonize the data content from different GWAS genotyping platforms so that… Read more »
With January officially in the bag, 2015 is off to a great start, especially for some of our customers who have recently published. I wanted to take a minute to share them with you. Sander van der Laan at University Medical Center Utrecht, published Variants in ALOX5, ALOX5AP and LTA4H are not associated with atherosclerotic plaque phenotypes: The Athero-Express Genomics Study which assessed the impact of common variants… Read more »
In recent months we have been updating our public annotation library to include the most recent versions of existing sources, as well as include new sources. All of these annotation sources are compatible with our three major products, VarSeq, SVS, and GenomeBrowse, and can be used for visualization, annotation, and filtering. dbNSFP NS Functional Predictions 2.8, GHI and dbNSFP Predictions… Read more »
One frequent question I hear from SVS customers is whether whole exome sequence data can be used for principal components analysis (PCA) and other applications in population genetics. The answer is, “yes, but you need to be cautious.” What does cautious mean? Let’s take a look at the 1000 Genomes project for some examples.
The Plant & Animal Genome XXIII Conference (PAG) was again a success. It’s the venue for leading genetic scientists and researchers involved in plant and animal research to meet with their peers. If anything the event continues to grow. The largest population of registrations tend to be from an Academic background (64%), with Industry (25%) and Government (11%) sectors comprising… Read more »
In 1914 the German cytologist Theodor Boveri coined the phrase “Cancer is a disease of the genome”. At this time his ideas were equally revolutionary as they were highly contested. Fast forward. More than hundred years later, Next-Generation Sequencing effectively permits a highly sensitive analysis of cancer cells. It can help us to understand mutations associated with cancer development and… Read more »
The Integrative Therapies Institute is soon hosting the annual, ITI 2015 conference January 23rd through the 25th in sunny San Diego and our own Dr. Andreas Scherer has been invited to speak. Some of the most prominent genomic and integrative medicine specialists will gather at ITI 2015 to share case studies and protocols with the community. Attendees can expect to… Read more »
If you haven’t been closely watching the twittersphere or other headline sources of the genetics community, you may have missed the recent chatter about the whole genome sequencing of 17 supercentenarians (people who live beyond 110 years). While genetics only explains 20-30% of the longevity of those with average life-spans, it turns out there is a number of good reasons… Read more »
There is a lot we can be grateful for at Golden Helix. The past year was marked by two major breakthrough launches. Earlier in 2014, we shipped SVS 8 which unified SVS with our GenomeBrowse product. We were able to improve SVS’ data management and visualization capabilities. In addition we added a number of new methods in SVS, such as SKAT-O, MM-KBAC, and various genomic prediction algorithms.
Once again, we will be kicking off our year with our annual trip to San Diego for PAG XXIII. This year, it could not come at a better time. Over the last few weeks, it has been bitter cold in Montana with temps barely reaching above zero degrees and I for one am looking forward to the warm sun. And… Read more »
It’s cliche, I know, but wow…2014 flew by! And what a great year it was for the Golden Helix team – we made upgrades to both GenomeBrowse and SVS and released a brand new product – VarSeq! In April, we released GenomeBrowse 2.0, which was a reflection of our most frequent user requests. Users now have the ability to upload… Read more »
Last month, I announced our Golden Helix Gives Back Campaign. During times like this, when funding is tight, we wanted to make a statement to our community. We at Golden Helix are committed to empower researchers and practitioners in the life science field. For those hard working people it is nice to catch a break from time to time. After… Read more »