In our latest VarSeq release, we updated our PhoRank algorithm with the ability to specify OMIM phenotype terms not present in HPO, as well as a general update to the algorithm to improve the results. In this post, we review the fundamentals of how PhoRank determines the ranking of genes in your VarSeq projects based on your input phenotype terms… Read more »
We are pleased to announce our next webcast, CNV Analysis in VarSeq – A User’s Perspective. The live event is is scheduled for Wednesday, April 19th at noon EST. Here are the specifics: Wednesday, April 19th 12:00 pm EST Clinical labs must have the ability to go from a collection of samples to a professional report documenting a short list… Read more »
It may be possible to say that annotating a variant correctly and accurately against gene transcripts is the most important job of a variant annotation and interpretation tool. We take it very seriously at Golden Helix as we support VarSeq and its use by our customers in both research and clinical contexts. It has been a source of frustration that… Read more »
Join our upcoming webcast : Wednesday, April 5th 12:00 pm EST Dr. Reza Sailani is a Research Fellow in the Genetics department at Stanford University. To provide an overview of his research, Sailani will present on the following two recent studies he has conducted: Association of AHSG with alopecia and mental retardation (APMR) syndrome: Alopecia with mental retardation syndrome (APMR) is… Read more »
Today, I am happy to announce the introduction of VS-CNV for Gene Panels and Exomes. We have developed this capability in partnership with PreventionGenetics. PreventionGenetics will use VarSeq CNV for analysis of gene panels, and in the future for exome sequencing. The software gives PreventionGenetics the opportunity to conduct a comprehensive CNV analysis on NGS data, in many cases eliminating the need… Read more »
ACMG 2017 is just around the corner! We are halfway through March already and it’s just about time to head off to sunny and warm Phoenix, Arizona. While the temps have been mostly mild for the last few weeks in Montana, I bet those of you in the northeast are looking forward to your time in Phoenix! You will find… Read more »
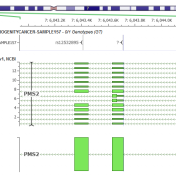
VarSeq will soon provide CNV exome analysis! In our webcast last week, we announced that we took our CNV caller, VS-CNV, to the next level. Along with the ability to call CNVs at the single-exon level in NGS gene panels, VarSeq can soon be used to call large loss of heterozygosity (LOH) and CNV events at the exome level. The combination… Read more »
In the past couple of weeks, the topic of the Filter and Quality fields in the popular ExAC population catalog has come up a number of times. It turns out that unlike the 1000 Genomes project, which decided to very heavily filter their variant list to only contain variants they consider high quality, ExAC chose to include more dubious variants… Read more »
This generation of scientists, clinicians and bioinformaticians have already elevated the standards for diagnosis, prediction and care, ultimately improving patient outcome for millions of people by leveraging genomic information. This trend is only going to continue. Next-gen sequencing has made its way into the clinic. Golden Helix supports the adoption of Precision Medicine by building products, such as our VarSeq… Read more »
Genome-wide association study (GWAS) technology has been a primary method for identifying the genes responsible for diseases and other traits for the past ten years. GWAS continues to be highly relevant as a scientific method. Over 2000 human GWAS reports now appear in scientific journals. In fact, we see its adoption increasing beyond the human-centric research into the world of… Read more »
Just as we expected, 2017 has kicked off with a flurry of new publications by our customers. We even had a publication from a client using our VarSeq software! Congratulations to all, please take a look at some of the articles we have highlighted below: Reza Sailani of Stanford University and colleagues published Association of AHSG with alopecia and mental retardation… Read more »
With a properly defined wet-lab and bioinformatics process, we are able to zero in on clinically relevant variants. How does a lab report on the outcome of their analysis? We find that most laboratories conduct their variant classification based on the guidelines formulated by the American College of Medical Genetics (ACMG) for inherited diseases. The ACMG guidelines for variant classification… Read more »
It may have been easy to miss in the drum-beat of monthly annotation updates we do here at Golden Helix, but there are a couple of things that are very special about the January update to the ClinVar database: We added new fields including HGVS names of variants and citations in PubMed for variants ClinVar nearly doubled in size by… Read more »
Acute lymphoblastic leukemia (ALL) is the most frequently diagnosed cancer in children and one of the leading causes of death due to disease in children. Dr. Daniel Sinnett, along with Pascal St-Onge and their colleagues at Sainte-Justine University Health center have been investigating the molecular determinants of the disease to improve detection, diagnosis and treatment. One particular area of study… Read more »
It sure is feeling like Christmas time in Montana with the piles of fluffy snow and negative temperatures! We are wrapping up the month with a few more publications from our clients, and we couldn’t be happier with how many articles were published in 2016! Congratulations to everyone who was able to get it done this year, and we are looking… Read more »
December’s webcast will provide the Golden Helix community with a more in-depth look at CNV analysis in VarSeq. On December 7th, Dr. Nathan Fortier will discuss the challenges and metrics surrounding CNV detection and then demonstrate VarSeq’s new capability from VCF to clinical report. Wednesday, December 7th @ 12:00 PM, EST Numerous studies have documented the role of Copy Number Variations (CNVs)… Read more »
While clinical assessments of germline mutations have been collected in ClinVar under the stewardship of the NCBI and the collaborate effort of many testing labs, the same type of resource has been missing for mutations that could informal clinical care in Cancer. Or at least, that is what I thought until I started to work with CIViC. With the stewardship of… Read more »
This year we have added some very important features to our software including CNV calling in VarSeq and integrating our premium annotations into SVS. We have also made many improvements to our software’s performance so that you can handle growing datasets with relative ease. In an effort to thank our community for their support throughout the year, we are happy… Read more »
ASHG 2016 is in our rear mirror. Again, it was bigger and better than the previous year. The conference hosted over 9,000 visitors from 66 countries. This gave the event a level of vibrancy that was evenly matched by the wonderful ambiance of the city of Vancouver. Nestled in between the two conference centers was a little pier offering spectacular… Read more »
The new Annotate and Filter algorithm is now available with the release of SVS 8.6.0, see the release notes for full details on all new and updated features. To access this new functionality, you simply need to update your SVS installation to the new version. The update can be done by clicking the Update Available link at the bottom of… Read more »