We would like to thank everyone who participated in our 2021 t-shirt design competition. It was great to see the amount of creativity our community expressed and was certainly a tough decision to make! We are pleased to announce this year’s winners: First Place – Bruce Eng For over 35 years, Bruce Eng has partnered with businesses, educational institutions, and… Read more »
Welcome to the August edition of our customer publications blog post! Each month we spotlight a few recently published articles by our incredible Golden Helix customers. With users spanning both research and clinical spaces, the topics vary widely across many fields. This month, we will be highlighting VSClinical users and the guided workflow. Host Genetics and Antiviral Immune Responses in… Read more »
Join the Golden Helix team at this year’s ESHG 2021 Virtual Conference! We will be presenting two different talks on our different product solutions and fielding any questions you might have. VSClinical: a comprehensive NGS clinical solution The first talk, VSClinical: a comprehensive NGS clinical solution, will be on Sun, 29 August, 14:00-15:00. This will be moderated by Golden Helix… Read more »
In this month’s Customer Publication blog, I will highlight four studies that provided further insights into conditions that are typically identified in early childhood. As you will see as you read the summaries of each publication, both Golden Helix software platforms (VarSeq and SNP & Variation Suite or SVS for short) were instrumental in exploring the genetic factors that influence… Read more »
We are honored to be selected as MirrorReview’s 10 Innovative Biotechnology Solution Companies for 2021. This award would not be possible without the incredible support from our customers and partners. Thank you to our entire community! If you are interested in reading our story, and why we were selected for this award, you can access the publication here: https://www.mirrorreview.com/golden-helix-pioneering-bioinformatics-solutions/. “Biotechnology… Read more »
The Genome Aggregation Database (gnomAD) is a resource developed by an international coalition of investigators, with the goal of aggregating and harmonizing both exome and genome sequencing data from a wide variety of large-scale sequencing projects (1). We have covered this annotation in-depth in other blog posts, but this resource contains over 125,000 exome sequences and around 16,000 whole genome… Read more »
Could leveraging CNV analysis for whole exome or genome sequencing help provide answers for the undiagnosed? Roughly 4% of the world’s population shares one common desire – a diagnosis (1). Though it may seem counterintuitive, a diagnosis, no matter how grim, can provide relief, validation, the chance to make plans, and have a discernible sense of future for individuals and… Read more »
Golden Helix’s solutions are leveraged in a wide variety of scientific fields. As always, we are thrilled to showcase some of the newly published articles written by our customers. Here are a few that utilized VarSeq’s powerful scalability analyzing whole exome sequencing data. WES can identify variations in the protein-coding region of any gene. Since most known mutations that cause… Read more »
Next-gen sequencing (NGS) comprises many sophisticated steps that are often compressed into three major sections: library prep, sequencing, and data analysis. Obviously, the goal is to simplify each of these steps, but more often than not, there is a need for multiple tools to complete each one. Regarding the data analysis, Golden Helix seeks to provide simple yet comprehensive solutions… Read more »
VSPipeline is becoming a very popular tool among VarSeq users as it is essential for creating repeatable clinical workflows that can be executed in automated fashion. Since VSPipeline is a command-line tool, I think it would be helpful to discuss some of the best practices along with helpful tips for getting the most out of VSPipeline. Some of you may be less familiar with VSPipeline, so I want to cover how to set up the first run along with sharing the helpful tips as they arise. … Read more »
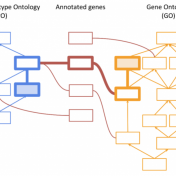
Thank you to those who attended our recent webcast, “PhoRank 2.0: Improved Phenotype-Based Gene Ranking in VarSeq”. For those who could not attend, you can find a link to the recording here. This webcast covered upcoming improvements to the PhoRank phenotype-based gene ranking algorithm based on literature published in the years since the algorithm’s development. The PhoRank Algorithm When performing… Read more »
A common feature request from Golden Helix customers is to curate and make available genome assemblies for different plant and animal species. These requests commonly come from SVS users as many research projects are being carried out, and having the genome assembly available for analysis is essential. That being said, Golden Helix has an SVS Tutorial available that walks users… Read more »
I learned about Batten disease from a childhood friend’s Facebook post. Over the course of a few months, her 8-year-old, Eva, the oldest of 4 daughters – Emily, Lucy, and Carly – was rapidly going blind. Baffled, doctors ran a genetic panel that returned a devastating result – the diagnosis of Juvenile Neuronal Ceroid Lipofuscinosis or Batten disease. A broad… Read more »
VSPipeline is a command-line interface that will provide high throughput environments the ability to tap the full power of VarSeq’s algorithms and flexible project template system from any command-line context, including the existing bioinformatics pipeline. This feature is a great resource for analyzing large sample volumes as it automates importing and annotating your data, which can help streamline your analysis… Read more »
In this month’s customer publication blog, both of our flagship software platforms are shown hard at work to cover many scientific investigational topics. You will get a glimpse of how VSClinical can be used to dive deeper into a specific gene associated with breast cancer and how SVS is enabling scientific discoveries in Agrigenomics and human health and wellness. Read… Read more »
Thanks for checking out this blog on how to to get more out of your VarSeq projects! The best way to show off one of my favorite new features that have been incorporated into the latest VarSeq 2.2.3 release is with a scenario: You work in a lab that processes and analyzes multiple whole exome samples for copy number variations,… Read more »
Golden Helix has just released VarSeq v2.2.3. In this update, there are notable changes that can improve CNV calling capabilities covered in this webcast. The topics discussed included: Accounting for GC content Improvements to CNV quality flags Using target filtering Updates to the CNV sensitivity and precision settings This blog post will elaborate on these capabilities and demonstrate how they… Read more »
Dr. Auber is the team leader for the molecular genetic diagnosis of hereditary diseases at the Institute for Human Genetics at Hannover Medical School (MHH). MHH is one of the largest hospitals in northern Germany, with one of the largest outpatient clinics for individuals and families dealing with hereditary cancer and predisposition syndromes. Dr. Auber was in high school when… Read more »
Thank you to those who attended the recent webcast, “High Precision Exome CNV Detection with VS-CNV”. For those who could not attend but wish to watch, here is a link to the recording. This webcast delved into the complex world of CNV calling for whole-exome samples, which presents unique challenges that require specific considerations and strategies. Over the past several months,… Read more »
Researchers and clinicians alike benefit from the powerful capabilities of Golden Helix’s software. Our tools are continuously validated, and we like to showcase a few articles each month that demonstrate the multitude of use cases and advancements in science. For our April installment, I would like to highlight the clinical space, with users spanning the globe, all with a common… Read more »