The adoption of genetic services is key to our ability to provide personalized medicine in the future. The goal is to better diagnose diseases, predict their outcomes, and to choose the best possible care option for a patient. Our part here at Golden Helix is to essentially build the equivalent of an MRI for the genome. In this process the latest… Read more »
Over the last decade, DNA sequencing has made vast technological improvements. With the cost of sequencing decreasing significantly, sequencing technology has become a product for the masses. The sequencing technology and programs that were once used exclusively by major research institutions are now becoming available in many research facilities around the globe. These tools produce large amounts of data sets… Read more »
Drumroll, please! Voting has come to a close for the 2014 Golden Helix T-shirt contest, and there were some clear favorites among the finalists. We are very excited at the wealth of creativity that came forward with this contest and are happy to announce that our final decisions have been made.
With the t-shirt submission deadline behind us, it’s time for the exciting part of the contest – picking the winners! We received a ton of fantastic designs and had a hard time narrowing them down. But, the Golden Helix team has picked seven designs that truly embody the Golden Helix spirit.
We released GenomeBrowse 2.0 earlier this year, allowing users to review all types of genomic data. Since then, it has received rave reviews from thousands of users around the world. Essentially, it’s the Google Earth app for genomic data. GenomeBrowse allows a user to sift through vast amounts of genomic data, and make it easy to focus on a single part… Read more »
On my flight back from this year’s Molecular Tri-Conference in San Francisco, I couldn’t help but ruminate over the intriguing talks, engaging round table discussions, and fabulous dinners with fellow speakers. And I kept returning to the topic of how we aggregate, share, and update data in the interest of understanding our genomes. Of course, there were many examples of… Read more »
Tis the season of quiet, productive hours. I’ve been spending a lot of mine thinking about file formats. Actually, I’ve been spending mine implementing a new one, but more on that later. File formats are amazingly important in big data science. In genomics, it is hard not to be awed by how successful the BAM file format is. I thought… Read more »
Genotype imputation is a common and useful practice that allows GWAS researchers to analyze untyped SNPs without the cost of genotyping millions of additional SNPs. In the Services Department at Golden Helix, we often perform imputation on client data, and we have our own software preferences for a variety of reasons. However, other imputation software packages have their own advantages… Read more »
Thanks to everyone for the great webcast yesterday. We had over 850 people register for the event and actually broke the record! Take that Bryce and Gabe! If you would like to see the recording, view it at: Mixed Models: How to Effectively Account for Inbreeding and Population Structure in GWAS. While preparing for this webcast, we chose to focus… Read more »
When researchers realized they needed a way to report genetic variants in scientific literature using a consistent format, the Human Genome Variation Society (HGVS) mutation nomenclature was developed and quickly became the standard method for describing sequence variations. Increasingly, HGVS nomenclature is being used to describe variants in genetic variant databases as well. There are some practical issues that researchers… Read more »
I’m a believer in the signal. Whole genomes and exomes have lots of signal. Man, is it cool to look at a pile-up and see a mutation as clear as day that you arrived at after filtering through hundreds of thousands or even millions of candidates. When these signals sit right in the genomic “sweet spot” of mappable regions with… Read more »
A few months ago, our CEO, Christophe Lambert, directed me toward an interesting commentary published in Nature Reviews Genetics by authors Bjarni J. Vilhjalmsson and Magnus Nordborg. Population structure is frequently cited as a major source of confounding in GWAS, but the authors of the article suggest that the problems often blamed on population structure actually result from the environment… Read more »
My investigation into my wife’s rare autoimmune disease I recently got invited to speak at the plenary session of AGBT about my experience in receiving and interpreting my Direct to Consumer (DTC) exomes. I’ve touched on this before in my post discussing my own exome and a caution for clinical labs setting up a GATK pipeline based on buggy variants… Read more »
In a recent GenomeWeb article by Tony Fong, “Sequenom’s CEO ‘Puzzled’ by Illumina’s Buy of Verinata, Lays out 2013 Goals at JP Morgan,” Harry Hixson, Sequenom’s CEO, expresses puzzlement over why its major supplier, Illumina, is acquiring a Sequenom competitor in Non-Invasive Prenatal Testing (NIPT), and thus apparently competing with one of its major customers. In a JP Morgan interview… Read more »
In preparation for a webcast I’ll be giving on Wednesday on my own exome, I’ve been spending more time with variant callers and the myriad of false-positives one has to wade through to get to interesting, or potentially significant, variants. So recently, I was happy to see a message in my inbox from the 23andMe exome team saying they had… Read more »
What prevents scientists from being more productive and if we knew, could we do anything about it? I’d like to look at an often overlooked, but huge productivity inhibitor — bad multitasking. Many people put “excellent multitasker” on their resume as a badge of honor. We laud the efficiency of a good multitasker — they are rarely idle — someone… Read more »
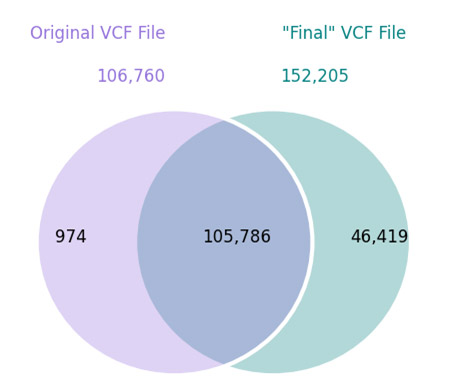
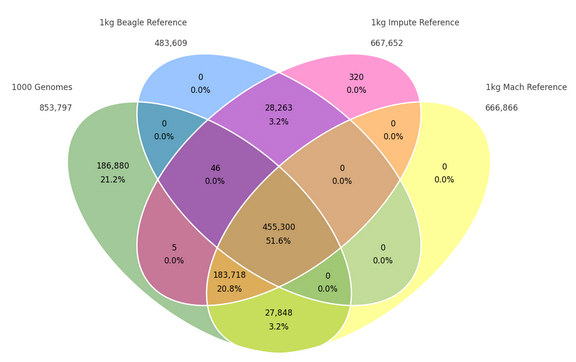
I recently curated the latest population frequency catalog from the 1000 Genomes Project onto our annotation servers, and I had very high hopes for this track. First of all, I applaud 1000 Genomes for the amount of effort they have put in to providing the community with the largest set of high-quality whole genome controls available. My high hopes are… Read more »
Type 2 Diabetes, Rheumatoid Arthritis, Obesity, Chrohn’s Diseases and Coronary Heart Disease are examples of common, chronic diseases that have a significant genetic component. It should be no surprise that these diseases have been the target of much genetic research. Yet over the past decade, the tools of our research efforts have failed to unravel the complete biological architecture of… Read more »
This month Biostatistics published online an open access article I co-authored with Dr. Laura Black from Montana State University: “Learning From Our GWAS Mistakes: From Experimental Design To Scientific Method.” The paper version is expected to come out in the April 2012 issue. I’m hoping that you will take the time to read it. And I’m hoping you will violently… Read more »
It’s that time again! We here at Golden Helix are excited to announce SVS 7.6 with more features for DNA-Seq analysis, the addition of RNA-Seq functionality, reorganization of SVS into “packages” including two new ones, and the release of new plot types enabled by Matplotlib. It’s certainly been busy as we pack all this into the sixth installment of the… Read more »