There used to be much energy expended at conferences, bioinformatics forums and even publications about what was the better strategy for interpreting variants of clinical significance: Rule-based filtering and classification mechanisms or rank-based prioritization through all-encompassing “pathogenicity” scores.
Both have shown to be effective.
Rule-based systems, as exemplified in this filtering diagram in Baylor’s ground-breaking paper on clinical whole-exome sequencing are easy to interpret and result in a clean separation of variants into buckets such as those defined by the ACMG guidelines.
Variant scoring and ranking systems have had successes and champions as well. In the early 2011 paper describing the novel Ogden syndrome, VAAST was used to pick out the candidate variants and ranked highly the one validated as causal (although the causal mutation was confirmed using filtering approaches using ANNOVAR and later Golden Helix as well).
As the years have passed, the pragmatic consensus seems to be: why not use both!
Now with VarSeq, you can.
VarSeq with CADD: Score Every Variant… Fast!
The University of Washington’s Combined Annotation Dependent Depletion (CADD) algorithm provides C-scores of “deleteriousness” for single nucleotide variants (SNV) as well as insertions/deletions (indels) in the human genome.
Since its publication and subsequent updates, it has been strongly leading the pack of algorithms that combine many genomic annotations into a single score meant to provide a relative pathogenicity for any variant.
The types of annotations integrated through their machine learning algorithm (SVM) range from conservation, population frequency, regulatory, functional and structural information. As a result, unlike the existing annotations in VarSeq such as dbNSFP, CADD provides scores for both coding and non-coding regions and for both SNVs and indels.
With the latest release of VarSeq, we have added a CADD annotation algorithm that can be licensed as an add-on to your VarSeq subscription. When enabled, you will be able to easily add CADD into your annotation, filtering and ranking VarSeq workflow.
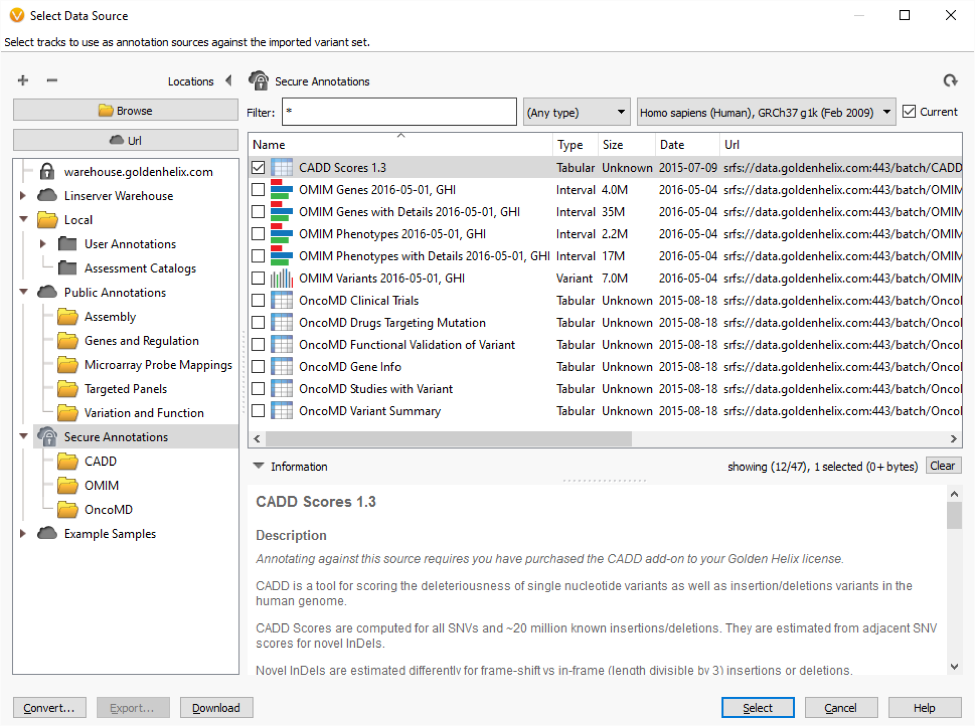 You can now select CADD Scores from our Secure Annotations repository. In a typical exome use case, 50K variants were annotated with CADD scores in under 3 minutes.
You can now select CADD Scores from our Secure Annotations repository. In a typical exome use case, 50K variants were annotated with CADD scores in under 3 minutes.
Like our OMIM and OncoMD, CADD will be listed in our Secure Annotation repository in VarSeq, and doesn’t require any downloads to start using.
And it’s FAST!
CADD scores are computed for every variant in your project at the rate of roughly 30,000 variants a minute.
Prioritize your Filtered Variants, or Aid Interpretation
The most likely use case for all those C-Scores is to either prioritize the candidate variants from your existing filtering pipeline or as an additional useful annotation to aid the interpretation process.
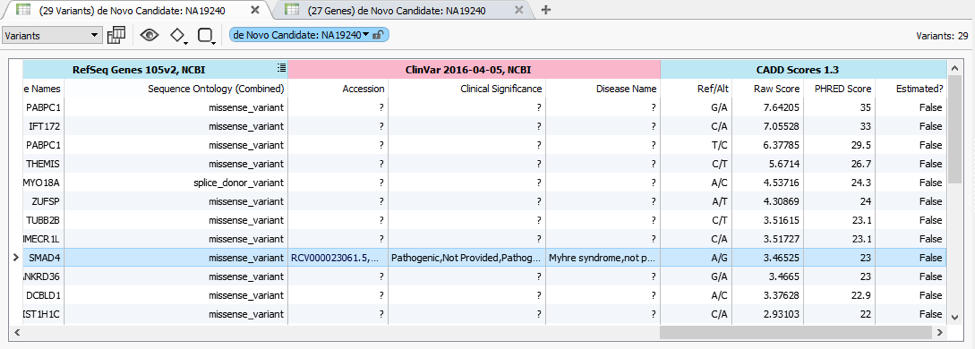 Sorting of the CADD scores here allows for the prioritization of these 29 candidate de Novo mutations in this trio. The SMAD4 variant is the one of interest and is in the top 10 candidates with no gene filtering.
Sorting of the CADD scores here allows for the prioritization of these 29 candidate de Novo mutations in this trio. The SMAD4 variant is the one of interest and is in the top 10 candidates with no gene filtering.
Similar to the function prediction voting we provide with dbNSFP, having the C-score available will give weight to the evidence for or against the causality of a variant to an individual’s symptoms or diagnosis.
The CADD scores in VarSeq will match exactly the ones provided for non-commercial use through the UW website, with one exception. While we can provide exact annotations for the precomputed 8.6 billion SNVs as well as a set of 20 million observed indels, “novel” indels not in this set are estimated and will be flagged by the “Estimated?” column in our annotation output.
These novel indels compose 1-5% of variants in a given dataset. Rather than have no score, we use a simple heuristic to combine the scores for the flanking or overlapping bases (see the track documentation for details) to capture the genomic context of the mutation.
These estimates correlate very highly with the exact scores (0.7 to 0.8 R^2 values, depending on the dataset) and thus rank exactly where you would expect them to be in a given analysis.
May All Your Variants Have Meaning
It’s very exciting to have CADD now fully integrated into VarSeq and available for your clinical variant interpretation and reporting workflows.
If you’re a VarSeq user, just ask your area manager for a trial of the CADD add-on. If you are thinking about checking VarSeq out, now is a better time than ever! You can request your evaluation here!