While clinical assessments of germline mutations have been collected in ClinVar under the stewardship of the NCBI and the collaborate effort of many testing labs, the same type of resource has been missing for mutations that could informal clinical care in Cancer.
Or at least, that is what I thought until I started to work with CIViC.
With the stewardship of Washington University in St Louis, an open source and iterative development model, an open access data licensing and a crowdsourcing but moderated wiki-like ability to submit, correct or comment on cancer mutations, it has the trajectory to become the ClinVar of Cancer.
We have just released our first curation of the CIViC monthly data releases and are planning on providing updates as we do with ClinVar at the beginning of each month.
Build with Purpose
While CIViC has some fantastic gene level information, we found the most applicable data to clinical labs is the Evidence Summaries.
Many users of VarSeq are providing genetic tests based on onco-gene panels, often designed to identify mutations that inform targeted molecular therapies, drug sensitivity and prognosis for specific cancers.
CIViC is a focused precision medicine resource for variants with published clinical evidence for the relationship between given mutations and diagnosis, prognosis, or response to a specific treatment of cancer.
Here are some of the key attributes captured by these Evidence Statements that are in our curated CIViC annotations as filterable, categorical fields:
- Evidence Type: What type of association exists between the evidence and a variant
- Predictive: Evidence of effect on therapeutic response
- Prognostic: Evidence of impact on disease progression, severity, or patient survival
- Diagnostic: Evidence of impact on patient diagnosis
- Predisposing: Evidence of effect on patient’s susceptibility
- Clinical Significance: The sub-type of evidence type that the statement presents
- Better Outcome: Demonstrates better than expected clinical outcome
- Poor Outcome: Demonstrates worse than expected outcome
- Positive: Associated with diagnosis of disease or subtype
- Negative: Associated with lack of disease or subtype
- Sensitivity: Evidence that pertains to a response to a therapeutic agent
- Resistance or Non-Response: Evidence that pertains to a lack of sensitivity to a therapeutic agent
- Evidence Level: The experimental method from which the evidence statement is derived
- A – Validated: Trusted association in clinical medicine that routinely informs treatment
- B – Clinical: Clinical evidence from clinical trials and other primary tumor data
- C – Case Study: Preclinical evidence from cell line studies, mouse models, and other in vitro or in vivo models
- D – Preclinical: Case study evidence from individual case reports in peer-reviewed journals
- E – Inferential: Inferential association made from experimental data
- Trust Rating: Trust Rating out of 5 Stars
- 1 out of 5 Stars: Claim is not supported well by experimental evidence
- 2 out of 5 Stars: Evidence is not well supported by experimental data, and little follow-up data is available
- 3 out of 5 Stars: Evidence is convincing, but not supported by a breadth of experiments
- 4 out of 5 Stars: Strong, well-supported evidence.
- 5 out of 5 Stars: Strong, well-supported evidence from a lab or journal with respected academic standing
While curating our annotation tracks, I also had the chance to experience the contributor side of the project by submitting various fixes to the representation of some variants with erroneously entered genomic coordinates or alleles. Each issue was quickly reviewed, sometimes with careful reproduction of my analysis, and the accepted changes integrated into the public database!
Similarly, on the open-source project side, we requested a change to allow for direct linking of the Evidence ID fields to facilitate URLs within VarSeq, and the feature was quickly added.
It Takes Two to Tango
While quite a few Evidence Statements in CIViC are about specific mutations with a list of alleles we can match like any other variant annotation source, quite a few are about mutations of larger genomic features, such as mutations of a codon, an exon or a gene.
For this reason, you will see two CIViC sources when you search for them in our Public Annotations repository:
The first one contains these Region based Evidence statements and will annotate all variants that overlap these regions. The second is the traditional variant annotation source with a Ref/Alt field that exactly matches your project mutations.
Apply CIViC, Filter, Interpret, Report
Let’s take a look at how CIViC adds to the cancer gene panel workflow we demonstrate in the VarSeq Cancer Gene Panel Tutorial.
To start, we open the VarSeq project, click Add -> Annotation, select the two CIViC tracks shown above and click Select. After browsing the resulting sources in our table, we decide to add a filter on the Clinical Significance column of each source to the filter chain:
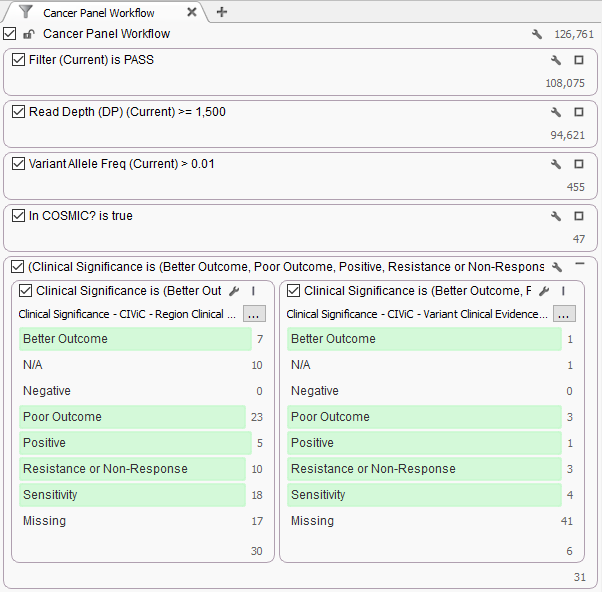
Quite a few of our COSMIC mutations were also found in CIViC. After drilling down on a few options, we find a mutation in KIT that is annotated against Exon 11 mutations in an Evidence Statement in the Melanoma samples we are investigating.
This is easy to see by plotting our project variants and adding the two CIViC sources as tracks to our GenomeBrowse view:
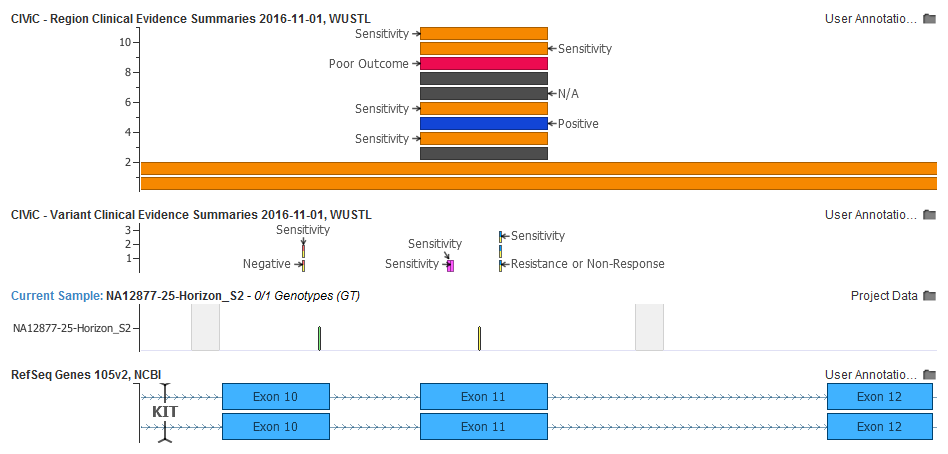
Here is the overlapping Evidence Statement related to Melanoma. Note you can follow the Evidence ID right to the corresponding record on the CIViC website.
| Variant Type | EXON 11 MUTATION |
| Variant Summary | c-KIT mutations in exon 11 lie within the juxtamembrane domain, and are very recurrent ingastrointestinal stromal tumors, often bearing a poorer prognosis than other KIT mutations. Cells harboring exon 11 mutations have shown sensitivity to the tyrosine kinase inhibitor imatinib, offering a better prognosis to patients treated with the drug in the first year. Small cohorts of melanoma patients harboring exon 11 KIT mutations have shown response to imatinib and sunitinib. |
| Disease | Melanoma |
| Drugs | Sunitinib |
| CIViC Evidence ID | 1224 |
| Evidence Type | Predictive |
| Clinical Significance | Sensitivity |
| Evidence Level | C – Case Study |
| Evidence Statement | 10 melanoma patients were evaluated after sunitinib treatment, among which 3 out of the 4 patients with KIT exon 11 mutations showed either complete or partial response. |
| PubMed ID | 22261812 |
| Citation | Minor et al., 2012, Clin. Cancer Res. |
| Trust Rating | 3 out of 5 Stars |
We can flag that variant for reporting and copy some of that info as supporting evidence!
On the Right Course
While there are other databases out there, such as MyCancerGenome that have similar goals as CIViC, the level of month-to-month activity and open and collaborative nature of the WUSTL folks puts it on the trajectory to be the dominant resource for clinical cancer genomics in our field.
We look forward to hearing from our users as they try out CIViC and hopefully contribute back their findings to this open access and contributor dependent resource.
Dear Gabe,
Great post. A nice summary of some of the key features of the CIViC interface. It is very helpful for us to get feedback from users and contributors who discover the CIViC on their own and have to “just figure it out” so to speak.
It was our hope in creating stable data releases, an open API, and open access licensing that it would help others to integrate CIViC into their own variant interpretation efforts. We are thrilled to have GoldenHelix as another early adopter!
Hey Malachi,
Thanks for the nice comment. And I really appreciate the open approach you guys have taken.
I attend the GA4GH calls and look forward to more resources being consolidated or cross-referenced by taking your lead.
Gabe