As VarSeq gains in popularity, we want to give Viewers and customers alike the opportunity to look at projects that are completed from start to finish. To this end, VarSeq (and VarSeq Viewer!) currently comes with two demonstration projects, Example TruSight Cardio Gene Panel and Example YRI Exome Trio Analysis. To access these projects from the VarSeq start page go to File > Open Example Project and if you’d like to get to the VarSeq Viewer please use the following link: http://goldenhelix.com/VarSeq/viewer-download.html. Once the viewer is downloaded the two projects will appear right away in the welcome window.
The TruSight Cardio Gene Panel contains two samples (NA12877 and NA12878) that were sequenced using the Illumina TruSight Cardio Sequencing Kit. Illumina makes the samples’ VCF and BAM files available for public download from their Illumina BaseSpace website. This gene panel provides comprehensive coverage of 174 genes with known associations to 17 inherited cardiac conditions, including cardiomyopathies, arrhythmias, aortopathies and more. The panel’s genes were expertly selected collaborating with researchers at the National Heart and Lung Institute at the Imperial College of London.
This example project uses a novel set of five filters that are different from the Hereditary Gene Panel Template that comes shipped with VarSeq. As per usual, the initial two filters are for quality assurance, selecting only those variants that pass the QC measures from the variant caller and a read depth filter.
The next filter comes from the ClinVar annotation source using the Clinical Significance category and allowing only those considered to be “Pathogenic” or of “Uncertain Significance” to pass through the filter card. This leaves just three variants for sample NA12878 and ten for sample NA12877.
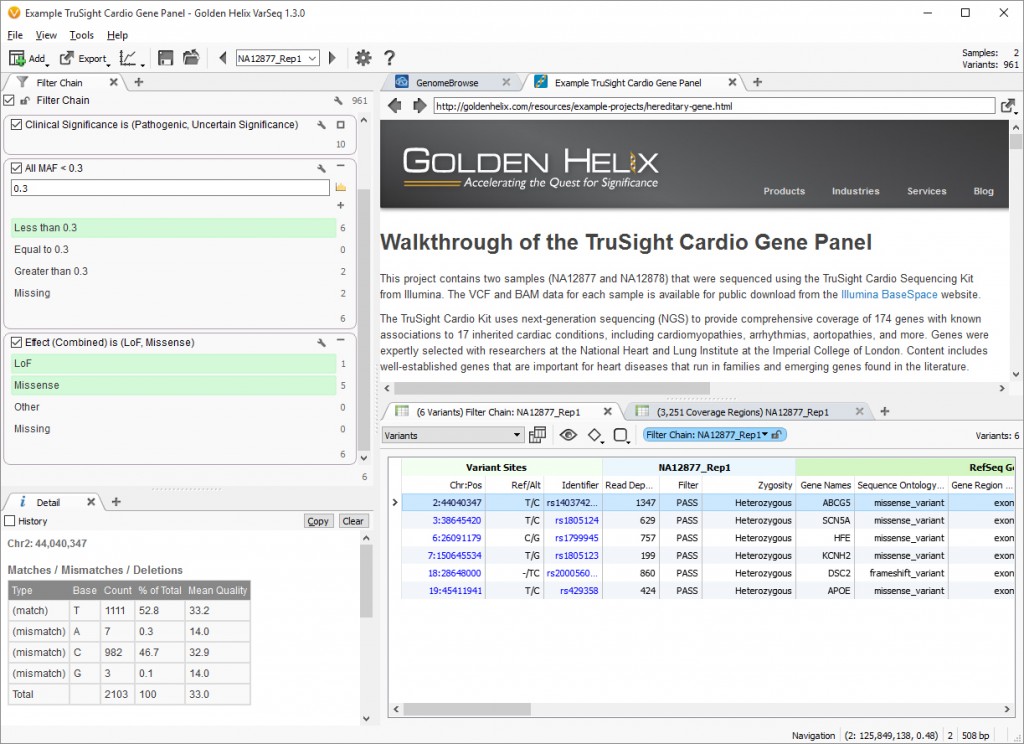
Then, the workflow narrows down based on minor allele frequency (MAF) based on the NHLBI Exome Variant Frequencies annotation source, and finally the last filter card identifies those variants that are considered to have a Loss of function (LoF) or to be a Missense variant according to the RefSeq Genes annotation source. This leaves sample NA12878 with a single variant to investigate and sample NA12877 with six variants (Fig. 1).
Fig. 1. Project view for Example Project TruSight Cardio Gene Panel. (Click image to enlarge)
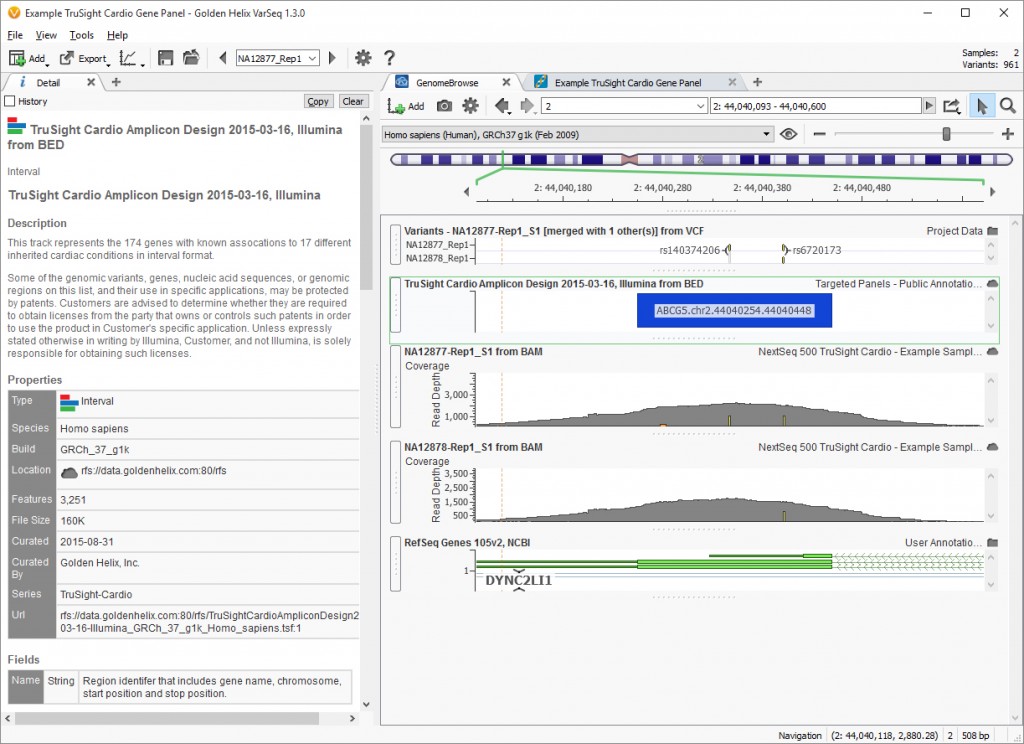
This is where having the associated BAM files can be helpful. The variants can be visualized in genomic space using the built-in software GenomeBrowse. You can see in the image below there is also a BED file. Having access to the sample BAM files also allows our customers to make use of the coverage statistics computation, another helpful QC measure!
Fig 2. Project view focusing on the GenomeBrowse view and the Detail view for the TruSight Cardio Amplicon Design BED file from Illumina. (Click image to enlarge)
The second example project analyzes a family trio from the Yoruban population from the HapMap Project (Mother-NA12938, Father-NA12939 and Child-NA12940). BAM files came from the 1000Genomes Phase3 Illumina Exome Alignment Project. We injected a de Novo variant into the daughter’s BAM file and then used SAMtools for variant calling on all three family members called simultaneously, creating a multi-sample VCF. The injected mutation is in the SMAD4 gene, which is associated with Myrhe Syndrome. This is found on chromosome 18, which for ease of use is the only chromosome included in the project instead of the whole exome.
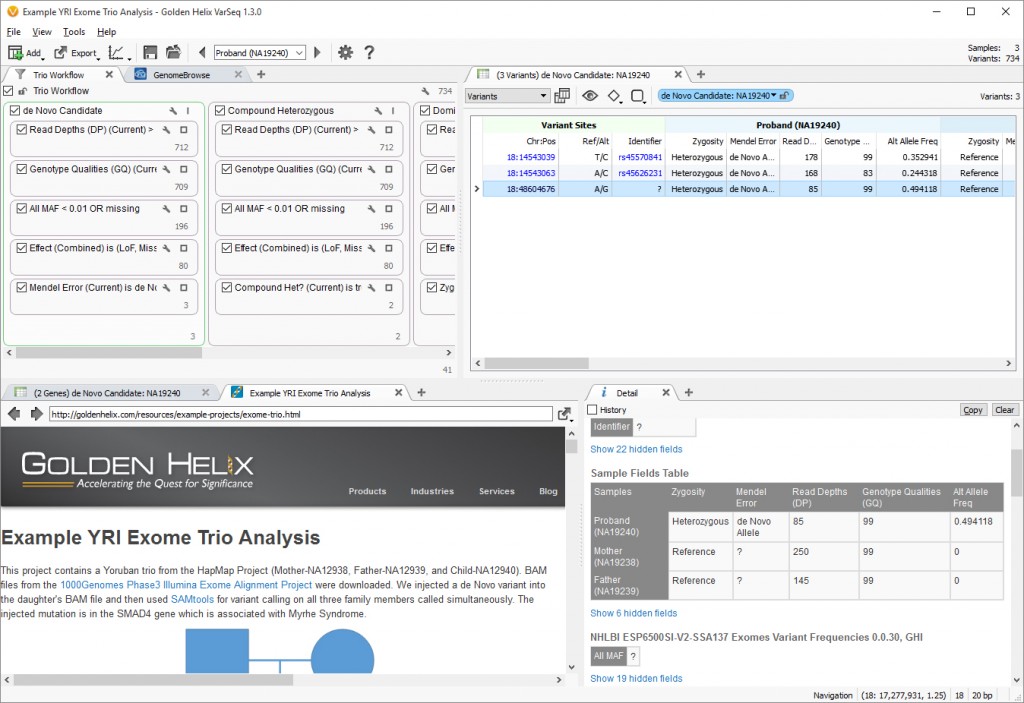
Fig 3. Project view for Example YRI Exome Trio Analysis. (Click image to enlarge)
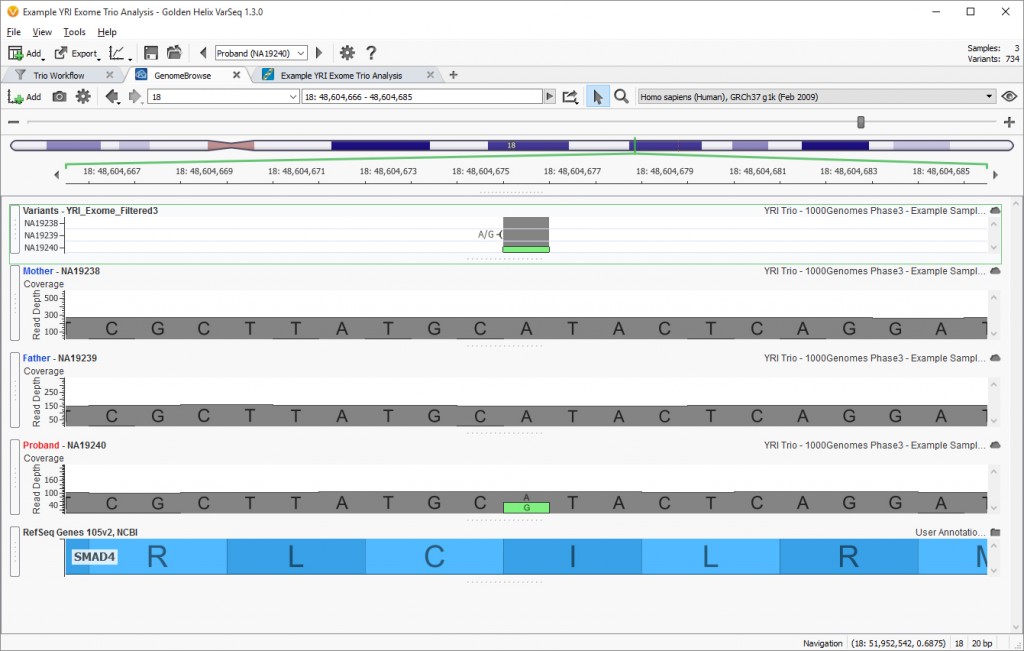
This project uses the Family Trio Template shipped with VarSeq and is set up with six different workflows analyzing different inheritance patterns. This is accomplished by changing the logic operator inside the filter view, switching it from “AND” to “OR” and then adding multiple filter containers to build each separate workflow. The union of each workflow is then seen at the bottom of the filter view for further analysis. In this project, there are 41 unique variants identified by the six workflows. Each of these workflows can be modified to further the analysis for customers interested in a particular workflow. In this case, we know the variant is novel to the proband, so looking at the De Novo Workflow we see there are only 3 variants from chromosome 18 that fit this inheritance pattern. The GenomeBrowse image below shows the causal variant present in the proband and missing from the mother and father VCF and BAM files (Fig. 2)
Fig 4. GenomeBrowse View of Family Trio VCF and BAM files. (Click image to enlarge).
This is just a summary of our current example projects, but each one also includes an HTML view explaining each filter chain and steps taken to come to conclusions in each project. These example projects offer our customers ideas for setting up filter chains for their own research and educating researchers through VarSeq Viewer on how to analyze different variant datasets. Finally we are happy to announce will be adding an additional example project analyzing a Tumor/Normal pair in the near future, so watch for this feature in our release notes!
If you have any questions about these example projects or want to get a copy of VarSeq or VarSeq Viewer, please contact support@goldenhelix.com.