The latest VarSeq release offers enhanced control over genome browse visualization computations. These computations, which are responsible for computing and caching the aggregate view of tracks within GenomeBrowse, are now more flexible. Typically, these tracks create files with the “.covtsf” extension and are used for zoomed-out density views of sources like VCFs and BAMs. Previously, the computation would commence automatically… Read more »
Calculating Residual Risk Residual risk is the risk remaining after a negative screening test. At the core of the calculation of residual risk resides Bayes’ Theorem. This theorem is used to calculate the probability of an outcome given the probability of events required for that outcome. The probability of the required events is compounded to calculate the combined probability of… Read more »
When interpreting fusions in their VCF format, it is not easy immediately grasp which side of the positions are adjacent in the resulting fusion. When interpreting and troubleshooting fusion variants, I usually find myself reaching for the VCF spec. If you, like me, are looking to speed up this process and gain a quick understanding of the fusions in your… Read more »
A new VSPipeline command, set_data_folder_path, designed to bolster consistent input usage. By introducing this innovative command, we aim to empower users with improved data organization, flexibility, and standardization for their clinical cases and analyses. Embracing this command will not only support reproducibility but also ensure accountability, ultimately paving the way for better-informed patient care decisions. Managing Annotations and References in… Read more »
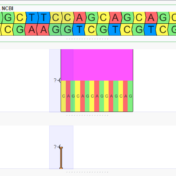
Discover how soft clip visualization can help you identify structural variants in your research and improve the accuracy of your findings. Soft clipping is a common technique in sequence alignment used to remove bases from the ends of reads that do not align with the reference sequence. Removing these bases typically improves alignment accuracy. However, when multiple reads are soft… Read more »
Unlocking the Potential of CRAM Files: The New VarSeq 2.3.0 Release for Enhanced Plotting, Coverage Analysis, and CNV Detection The CRAM (Compressed Reference-oriented Alignment Map) file format was conceived in 2011 as a more space-efficient way to store alignment data. It saves space over the previous standard BAM (Binary Alignment Map) by only storing the differences between each read and… Read more »
Merging variant records, VCFs, across samples is important when performing trio or family analysis as it ensures that hereditary relationships can be properly inferred. There are many ways to represent a single variant. Insertions and deletions may be right or left aligned, prefixes and suffixes can be added, and adjacent variants in the same sample may be combined or split… Read more »
In VarSeq 2.2.1, you can set template annotation sources to automatically update to the latest version. Previously, VarSeq templates were frozen in time. Now, each new project created from a template would use the same source that was used when the template was created. When you save a template, you can have the sources automatically update to the latest version…. Read more »
In the previous two articles, we explored the different steps of a clinical workflow. The first post covered the automated analysis that creates a VarSeq project. While the second post covered the interpretation steps and generation of a clinical report. These posts illustrated the ease with which these complex tasks can be carried out. Today we’ll dig a little bit… Read more »
In the previous blog post, we covered the automated steps to create a VarSeq project. Today we will examine the active analysis steps. These are the steps that require human interpretation to analyze the clinically relevant variants. A lab tech can take the first pass at the output in the generated VarSeq project. They can perform the quality control and… Read more »
Automating a clinical workflow creates a stable and repeatable clinical analysis. Automation reduces the potential to introduce human error, helps in regulatory compliance, and improves the precision of the clinical results. It is important to know that if you run a sample through your clinical pipeline, you are going to get the same results today as you will in 6… Read more »
The power of VSPipeline is in it’s ability to automate VarSeq workflows. Using VarSeq to create a pipeline template is great because it allows you to dial in the applied filters as well as interactively organize the annotations and applied algorithms. Automating a workflow with VSPipeline is straightforward when beginning with an existing project. However, there are several steps that… Read more »
As VarSeq continues its adoption amongst clinical labs and researchers looking for reproducible workflows for variant annotation, filtering and interpretation, we have continued to prioritize the addition of features to assess the quality of the upstream data at a variant, coverage and now sample level. The Importance of Quality Assurance Sample prep and sequencing problems are difficult to detect through the analysis… Read more »